Machine learning assisted methods for the identification of low toxicity inhibitors of Enoyl-Acyl Carrier Protein Reductase (InhA)
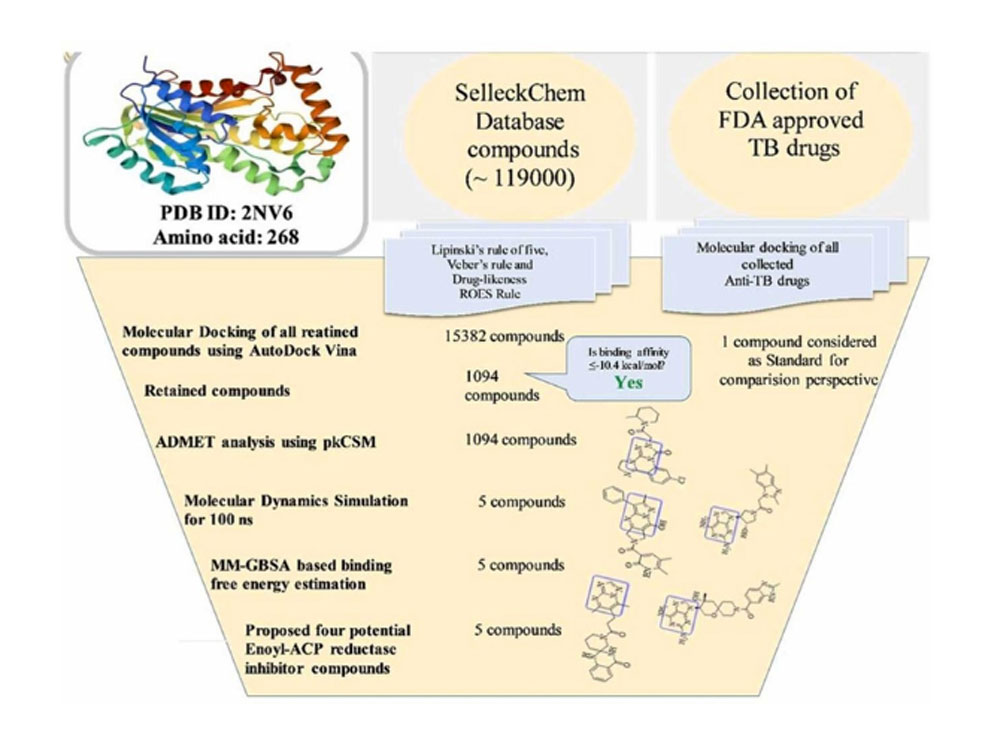
Tuberculosis, a deadly disease with ancient roots, continues to pose a global health threat. We have employed artificial intelligence and machine learning integrated approaches to identify potential new treatments by targeting the InhA protein in the tuberculosis-causing bacteria. A large chemical database ‘SelleckChem’ was meticulously filtered using advance computational methods to pinpoint molecules with promising drug-like properties. Molecular dynamics simulations and machine learning based ADMET analyses confirmed the potential of five identified compounds to effectively bind to the InhA protein and disrupt its function. Crucially, these molecules were predicted to be safe and easily producible, offering hope for the development of new anti-tuberculosis drugs. Article “Machine Learning Assisted Methods for the Identification of Low toxicity Inhibitors of Enoyl-Acyl Carrier Protein Reductase (InhA)” can check out in Science Direct.

Leave a Reply
Want to join the discussion?Feel free to contribute!