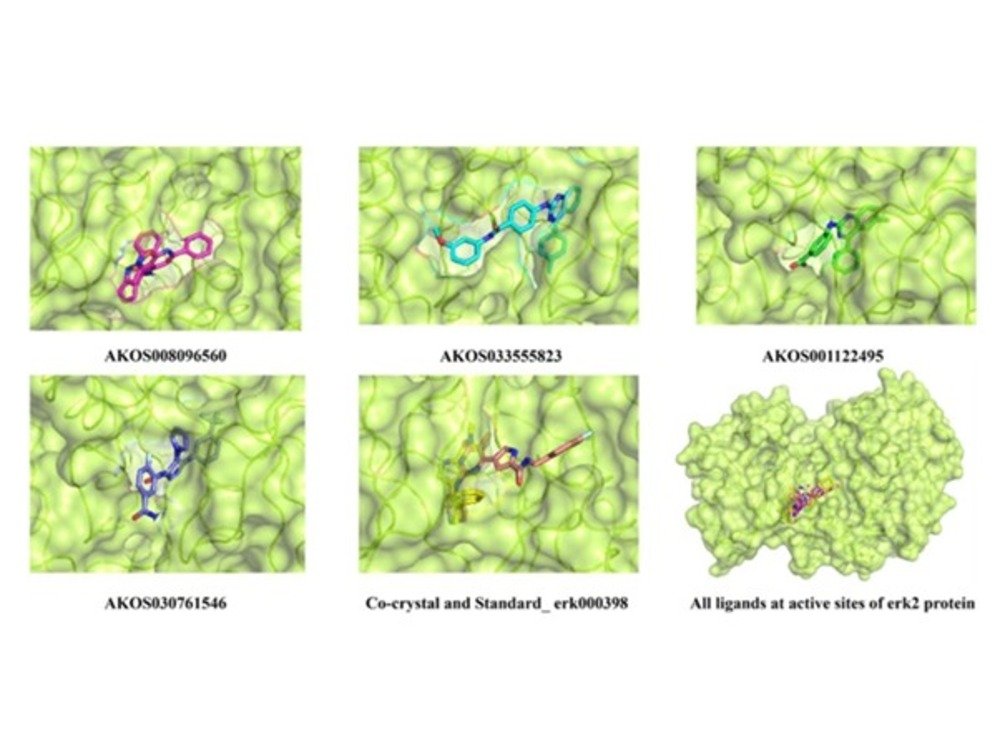
Machine learning-integrated and fingerprint-based similarity search against the immuno-oncology library for identification of novel ERK2 inhibitors
The extracellular signal-regulated kinase 2 (ERK2) protein plays a pivotal role in regulating cell division cycles and signal ing pathways essential for various biological processes. ERK2 inhibition is a promising therapeutic approach for diseases like cardiovascular deformities, neurodegenerative disorders, and other forms of cancers. The current study presents novel compounds potentially inhibiting ERK2 activity, thus disrupting its cellular functions. A thorough structural assessment of the available crystallographic information was undertaken. The protein’s active site was deciphered, and the experimental grid space of inhibitors interaction was allocated. The study proceeded further with a precise inhibitor search employing a “similarity search” algorithm based on the previously reported kinase inhibitors. Schematic virtual screening method combined with molecular docking steps were executed to enlist the probable hits. AI/ML-based pharmacokinetics proper ties helped streamline hits’ initial chemical space and select the most potent leads. Complexes formed by these compounds were analyzed for their stability by molecular dynamics (MD) simulations. Post dynamics statistical calculations, viz., pro tein backbone and ligand RMSD, the radius of gyration, and the constitutive amino acids fluctuations (RMSF), confirmed the protein–ligand association over a period of 300 ns. The magnitude of co-ordinations was estimated by intermolecular H-bond count and the MMGBSA calculations. The free energy landscape (FEL) and principal component analysis (PCA) demonstrated the thermodynamical feasibility of the complex formation with an affinity greater than the previously reported inhibitors. This study, thus, presents a promising avenue for advancing the drug discovery process by identifying novel ERK2 protein inhibitors with potential benefits for healthcare. Check out this published article, “Machine learning-integrated and fingerprint-based similarity search against the immuno-oncology library for identification of novel ERK2 inhibitors” in Structural Chemistry


Leave a Reply
Want to join the discussion?Feel free to contribute!