Molecular Simulations and Machine Learning Methods for the Identification of Novel Aurora A Kinase Inhibitors
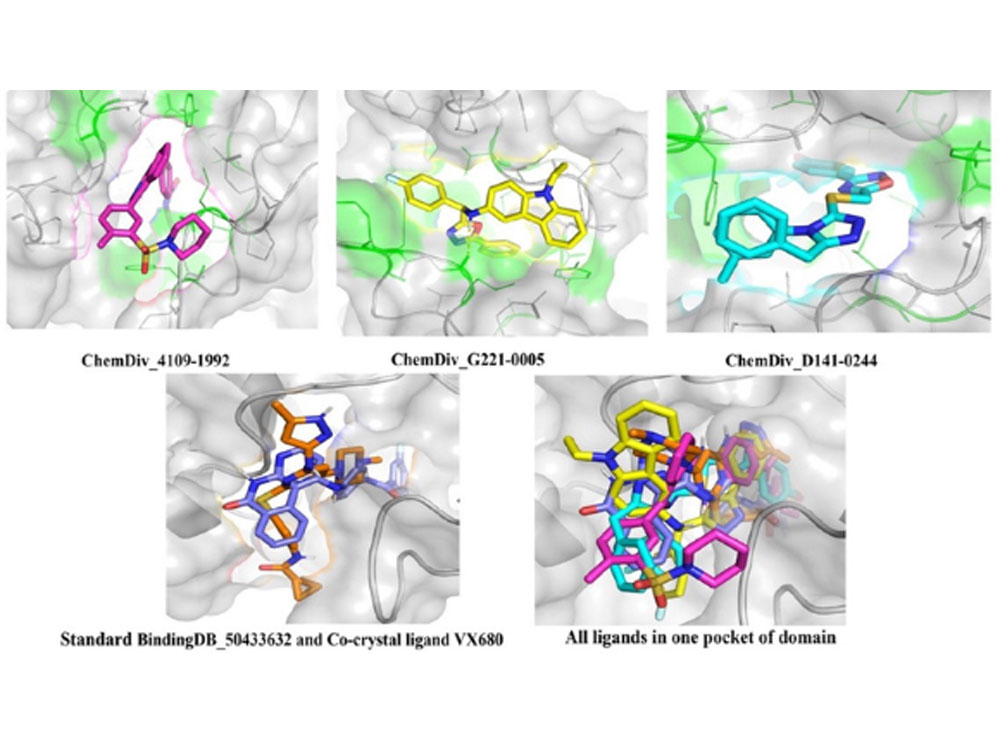
Aurora kinase A (AAK) is a critical regulator of mitosis, the process of cell division. It plays essential roles in cell cycle regulation, centrosome maturation, spindle assembly, and chromosome segregation, all crucial for accurate daughter cell formation. Deregulation of AAK expression and activity has been linked to various human diseases, particularly cancer, where increased AAK expression or activity contributes to cancer development and progression. In our recent study, we have employed artificial intelligence and machine learning techniques to identify potential inhibitors of AAK. Particularly we used multi-step molecular docking via AutoDock Vina and PLANTS to screen a ChemDiv kinase-specific inhibitor library against AAK. The study successfully identified three hit compounds with perfect binding in the active site pockets of AAK, comparable to the standard BindingDB compound and the co-crystal ligand VX-680 binding mode. These findings underscore the potential of computational drug discovery, bolstered by AI and machine learning, in identifying promising AAK inhibitors for improved cancer management. Published research article “Molecular Simulations and Machine Learning Methods for the Identification of Novel Aurora A Kinase Inhibitors” available in Journal of Biomolecular Structure & Dynamics.

Leave a Reply
Want to join the discussion?Feel free to contribute!